Phycodnaviridae
| Phycodnaviridae | |
|---|---|
| Virus classification | |
| (unranked): | Virus |
| Realm: | Varidnaviria |
| Kingdom: | Bamfordvirae |
| Phylum: | Nucleocytoviricota |
| Class: | Megaviricetes |
| Order: | Algavirales |
| Family: | Phycodnaviridae |
| Genera | |
Phycodnaviridae is a family of large (100–560 kb) double-stranded DNA viruses that infect marine or freshwater eukaryotic algae. Viruses within this family have a similar morphology, with an icosahedral capsid (polyhedron with 20 faces). As of 2014, there were 33 species in this family, divided among 6 genera.[1][2] This family belongs to a super-group of large viruses known as nucleocytoplasmic large DNA viruses. Evidence was published in 2014 suggesting that specific strains of Phycodnaviridae might infect humans rather than just algal species, as was previously believed.[3] Most genera under this family enter the host cell by cell receptor endocytosis and replicate in the nucleus. Phycodnaviridae play important ecological roles by regulating the growth and productivity of their algal hosts. Algal species such Heterosigma akashiwo and the genus Chrysochromulina can form dense blooms which can be damaging to fisheries, resulting in losses in the aquaculture industry.[4] Heterosigma akashiwo virus (HaV) has been suggested for use as a microbial agent to prevent the recurrence of toxic red tides produced by this algal species.[5] Phycodnaviridae cause death and lysis of freshwater and marine algal species, liberating organic carbon, nitrogen and phosphorus into the water, providing nutrients for the microbial loop.[6]
Taxonomy
[edit]Group: double-stranded DNA
- Family: Phycodnaviridae
- Genus: Chlorovirus
- Acanthocystis turfacea chlorella virus 1
- Hydra viridis Chlorella virus 1
- Paramecium bursaria Chlorella virus 1
- Paramecium bursaria Chlorella virus A1
- Paramecium bursaria Chlorella virus AL1A
- Paramecium bursaria Chlorella virus AL2A
- Paramecium bursaria Chlorella virus BJ2C
- Paramecium bursaria Chlorella virus CA4A
- Paramecium bursaria Chlorella virus CA4B
- Paramecium bursaria Chlorella virus IL3A
- Paramecium bursaria Chlorella virus NC1A
- Paramecium bursaria Chlorella virus NE8A
- Paramecium bursaria Chlorella virus NY2A
- Paramecium bursaria Chlorella virus NYs1
- Paramecium bursaria Chlorella virus SC1A
- Paramecium bursaria Chlorella virus XY6E
- Paramecium bursaria Chlorella virus XZ3A
- Paramecium bursaria Chlorella virus XZ4A
- Paramecium bursaria Chlorella virus XZ4C
The taxonomy of this family was initially based on host range: chloroviruses infect chlorella-like green algae from freshwaters; whereas, members of the other five genera infect marine microalgae and a some species of brown macroalgae. This was subsequently confirmed by analysis of their B-family DNA polymerases, which indicated that members of the Phycodnaviridae are more closely related to one another, in comparison to other double stranded DNA viruses, forming a monophyletic group.[7][8][9] The phycodnaviruses contain six genera: Coccolithovirus, Chlorovirus, Phaeovirus, Prasinovirus, Prymnesiovirus and Raphidovirus. The genera can be distinguished from one another by, for example, differences in life cycle and gene content.[8]
Structure
[edit]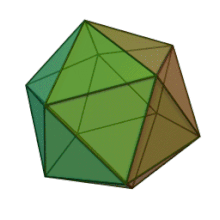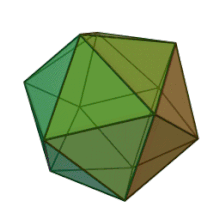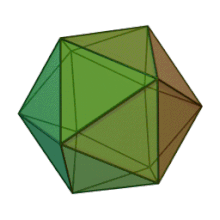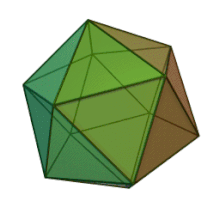
All six genera in the family Phycodnaviridae have similar virion structure and morphology. They are large virions that can range between 100 and 220 nm in diameter. They have a double-stranded DNA genome, and a protein core surrounded by a lipid bilayer and an icosahedral capsid.[10] The capsid has 2, 3 and 5 fold axis of symmetry with 20 equilateral triangle faces composing of protein subunits. In all known members of the Phycodnaviridae the capsid is composed of ordered substructures with 20 trisymmetrons and 12 pentasymmetrons made up of donut-shaped trimeric capsomers, where each capsomer is made up of three monomers of the major capsid protein. If all the trimeric capsomers are identical in structure, the virion capsid contains 5040 copies of the major capsid protein in total with a triangulation number of 169. At the five-fold vertices there are 12 pentamer-capsomers consist of different proteins. The protein(s) that can be found below the axial channel of each pentamer may be responsible for digesting the host cell wall during viral infection. The species Phaeocystis puchetii virus from the genus Prymnesiovirus has the largest capsid structure in the Phycodnaviridae family.[11]
The lipid bilayer membrane in phycodnaviruses is not well understood or researched. Some studies suggested that the membrane originates from the endoplasmic reticulum and may also be directly acquired from the host cell membrane during viral assembly. Although members of the family Phycodnaviridae are highly diverse, they share very conserved genes involved with virion morphology or structure.[citation needed] Despite the similarity of the capsid structure of phycodnaviruses, recent experiments have identified morphological differences among members in this family. Emiliania huxleyi virus 86 (EhV-86), a coccolithovirus strain, differs from its algal virus counterparts in that its capsid is enveloped by a lipid membrane.[12] In addition, recent 3D reconstruction experiments revealed that the chlorella virus PBCV-1 has a 250A-long cylindrical spike extending from one of its vertices. EhV-86 may also possess a spike or tail structure.[13]
Genome
[edit]Phycodnaviruses are known for their large double-stranded DNA genomes ranging from 100kb to over 550 kb with 40% to 50% GC content.[8] Currently, complete genome sequences are available for several members of the family Phycodnaviridae (including six chloroviruses, two phaeoviruses, several prasinoviruses and a coccolithovirus) and there are also some partial sequences available for a different coccolithovirus.[14][15][16][17]
The genome structures of phycodnaviruses have considerable variation. The chlorovirus PBCV-1 has a linear 330 kb genome with non-permuted double-stranded DNA that is covalently closed by hairpin termini. Similarly, the EsV-1 phaeovirus has a linear double-stranded DNA genome with inverted repeats that have almost perfect homology. These inverted repeats could facilitate effective circularization of the genome and for a period of time it has been suspected that EsV-1 has a circular genome.[15] The EhV-86 coccolithovirus is suggested to have both linear and circular genomes at different phases during DNA packaging. PCR amplification reveals random A/T overhangs, detection of DNA ligases and endonucleases hinting that a linear genome may be packaged and circularizes during DNA replication.[16][18] The phycodnaviruses have compact genomes for replication efficiency with approximately one gene per 900 to 1000 bp of genome sequences.[16] The EsV-1 phaeovirus is an exception with 231 protein encoding genes, which means it has one gene per approximately 1450 bp. In spite of the compact genomes typically found in viruses, Phycodnaviridae genomes have repetitive regions usually near the terminal ends and certain tandem repeats located throughout the genome. It is suggested that these repetitive sequences may play a role in gene recombination that allows the virus to exchange genetic information with other viruses or the host cell.[19]
Phylogeny
[edit]![The evolutionary history was inferred by using the Maximum Likelihood method based on the JTT matrix-based model [1]. The bootstrap consensus tree inferred from 100 replicates is taken to represent the evolutionary history of the taxa analyzed. Branches corresponding to partitions reproduced in less than 50% bootstrap replicates are collapsed. The percentage of replicate trees in which the associated taxa clustered together in the bootstrap test (100 replicates) are shown by the size of red node on each brach. Initial tree(s) for the heuristic search were obtained automatically by applying Neighbor-Join and BioNJ algorithms to a matrix of pairwise distances estimated using a JTT model, and then selecting the topology with superior log likelihood value. The analysis involved 26 amino acid sequences. There were a total of 2599 positions in the final dataset. Evolutionary analyses were conducted in MEGA7.](http://upload.wikimedia.org/wikipedia/commons/thumb/4/48/Molecular_Phylogenetic_analysis_of_NCLDV_members_by_Maximum_Likelihood_method.png/274px-Molecular_Phylogenetic_analysis_of_NCLDV_members_by_Maximum_Likelihood_method.png)
Viruses belonging to Phycodnaviridae harbor double-stranded DNA genomes with sizes of several 100kbp, which together with other Megavirales (e.g. Iridoviridae, Pandoraviridae and Mimiviridae) are named nucleocytoplasmic large DNA viruses. Because of their large genome sizes and various proteins that are encoded, viruses of Phycodnaviridae are challenging the traditional concepts that viruses are small and simple "organisms at the edge of life". Phylogenetic analyses of core genes based on gene concatenation,[21] individual phylogenies of the DNA polymerase,[22] and the major capsid protein,[23] indicate the close evolutionary relationships among members of Phycodnaviridae and between Phycodnaviridae and other families of nucleocytoplasmic large DNA viruses.
Life cycle
[edit]| Genus | Host details | Tissue tropism | Entry details | Release details | Replication site | Assembly site | Transmission |
|---|---|---|---|---|---|---|---|
| Raphidovirus[24] | Alga | None | Cell receptor endocytosis | Lysis | Nucleus | Cytoplasm | Passive diffusion |
| Coccolithovirus | Alga | None | Cell receptor endocytosis | Budding | Nucleus | Cytoplasm | Passive diffusion |
| Phaeovirus | Alga | None | Cell receptor endocytosis | Lysis | Nucleus | Cytoplasm | Passive diffusion |
| Chlorovirus | Alga | None | Cell receptor endocytosis | Lysis | Nucleus | Cytoplasm | Unknown |
| Prymnesiovirus | Alga | None | Cell receptor endocytosis | Lysis | Nucleus | Cytoplasm | Passive diffusion |
| Prasinovirus | Alga | None | Cell receptor endocytosis | Lysis and budding | Nucleus | Cytoplasm | Passive diffusion |
Raphidovirus
[edit]In Raphidovirus (likely misspelled Rhaphidovirus), there is only one species, Heterosigma akashiwo virus (HaV), which infects the unicellular alga, Heterosigma akashiwo. H. akashiwo is a member of the class Raphidophyceae, a bloom forming species and is widely distributed in temperate and neritic waters.[21] Several other types of viruses infecting H. akashiwo have been isolated and are not to be confused with HaV, such as the H. akashiwo RNA virus (HaRNAV).[25] and H. akashiwo nuclear inclusion virus (HaNIV).[26][4] As HaV was first isolated and characterized in 1997,[4] information about the life cycle is limited.
HaV specifically infects H. akashiwo and does not infect other marine phytoplankton species tested.[4] The mechanisms determining the virus-host specificity is not well understood. Tomaru et al. (2008)[4] suggest that virus-host specificity maybe caused by unique interactions between a viral ligand and a host receptor. In a study by Nagaski et al., virus particles were found inside the host cytoplasm at 24 hours post-infection. The latent period or lysogenic cycle was estimated to be 30–33 h with an average burst size (number of viruses produced after lysis) of 770 per cell. Virus particles were found in the subsurface area and in the viroplasm area[5]
Coccolithovirus
[edit]
In 2009, MacKinder et al. elucidated the entry mechanism of the genera Coccolithovirus.[12] Using confocal and electron microscopy, the researchers demonstrated that the virus strain EhV-86 uses a unique infection mechanism, which differs from other algal viruses, and shows a greater similarity to the entry and exit strategies seen in animal-like nucleocytoplasmic large double stranded DNA viruses (nucleocytoplasmic large DNA viruses). EhV-86 differs from its algal counterparts in that its capsid is enveloped by a lipid membrane. EhV-86 enters cells by endocytosis (the process by which food or liquid particles are taken into the cell by a vesicle), or direct fusion (the viral envelope fuses with the host membrane). EhV-86 entry by endocytosis results in an additional membrane coat surrounding the capsid encapsulated genome. Regardless of the mechanism of entry, the capsid enters the cytoplasm intact. After entering the cell, the viral capsid disassembles and the DNA is released into the host cytoplasm or directly into the nucleus. EhV-86 is unique to other phycodnaviruses as it encodes six RNA polymerase subunits. Neither PBCV-1 nor ESV-1, for example encodes RNA polymerase components.[8] Viral RNA polymerase genes are not transcribed until at least 2 hours post infection (p.i). At 3–4 p.i, virions are assembled in the cytoplasm, with the help of ATPase (a DNA packaging protein) and transported to the plasma membrane where they are released from the host via a budding mechanism. In this budding mechanism, EhV-86 gains an outer membrane from the host membrane.[12] Burst size ranges from 400 to 1000 particles per cell.[8]
A cluster of sphingolipid-producing genes have been identified in EhV-86. Researchers have found that the production of viral sphingolipids produced during the lytic stage are involved in programmed cell death in coccolithophore populations. A high correlation was found between glycosphingolipid (GSL) production and caspase activity during the lytic stage in infected cells. Caspases are a family of protease enzymes involved in programmed cell death. The researchers also found that a critical concentration of GSLs (>0.06 mg/ml) is required to initiate cell lysis. Thus, the authors suggest that the production of GSLs to a critical concentration may be part of a timing mechanism for the lytic cycle. The authors also suggest that these biomolecules may be able to induce programmed cell death in other unaffected cells, thus serving as an algal bloom termination signal.[27]
Phaeovirus
[edit]Coccolithoviruses and phaeoviruses have been described as having opposing life strategies. The coccolithovirus possesses an Acute life strategy characterized by high reproduction and mutation rates and greater dependency on dense host populations for transmission. Phaeoviruses possess a Persistent life strategy where infection may or may not cause disease, and the genome is passed from parent to offspring.[28]
Phaeoviruses infect the Ectocarpales brown algae, which is an order of filamentous brown algae. One of the most studied phaeoviruses is Ectocarpus siliculosus virus, most commonly known as EsV-1.[28] The EsV-1 virus only infects the single-celled gametes or spores of E. siliculosus. Vegetative cells are immune to infection, as they are protected by a rigid cell wall.[29] Following infection, one copy of the viral DNA is incorporated into the host genome. The EsV-1 viral genome is then replicated and virions are assembled in the sporangia or gametangia of infected plants.[30] Viruses are subsequently released via lysis of reproductive cells, stimulated by changes in environmental conditions, such as an increase in temperature.[31] In healthy plants, environmental stimuli synchronize the release of gametes and zoospores into the surrounding water.[31] Free virus particles can then re-infect free-swimming gametes or spores of healthy plants. Infected gametes or spores undergo mitosis, forming infected plants and all cells of the progeny plant contain viral DNA. However, viral particles are only produced in the reproductive cells of the algae, while viruses remain latent in vegetative cells. In infected sporophytes, cells undergo meiosis and produce haploid spores. The EsV genome is transmitted in a Mendelian manner, where half of the progeny contain viral DNA. Often algae from infected spores are indistinguishable from algae derived from healthy spores, but are partially or fully incapable of reproduction.[29][30]
Chlorovirus
[edit]Chloroviruses are the only viruses characterized thus far that infect freshwater algae.[32] The hosts of chloroviruses are zoochlorellae, which are endosymbiotic green algae commonly associated with hosts Paramecium bursaria, coelenterate Hydra viridis, or the heliozoan Acanthocystis turfacea.[33] In the ciliate Paramecium bursaria, for example, the algae lives within the cells of the host, providing nutrients via photosynthesis. Living inside the cells of the ciliate offers protection for the algae, and a mode of transportation. Zoochlorellae are resistant to infection in their symbiotic state. When the relationship between the algae and host is disrupted, for example, through grazing by copepods, infection by chloroviruses is permitted.[34]
The life cycle of the chlorovirus infecting Paramecium bursaria, known as PBCV-1 has been studied in detail [citation needed]. Cryo-electron microscopy and 3D reconstruction of the viral capsid shows that there is a long 'spike' structure which first contacts the cell wall and likely serves to puncture the cell wall of the host. The PBCV-1 virus is specific to its host and recognition is mediated by the interaction of virus surface proteins with algal surface carbohydrates. Following attachment of the virus to the host's cell wall, capsid-bound glycolytic enzymes break down the cell wall. The viral membrane likely fuses with the host membrane, allowing the viral DNA to enter the cytoplasm, leaving an empty capsid on the outside. As PBCV-1 lacks an RNA polymerase gene, the virus must use the host cell's machinery to produce viral RNA. Thus, the viral DNA quickly moves to the nucleus where early transcription is initiated 5–10 minutes post infection. Within minutes of infection, host chromosomal degradation occurs, inhibiting host transcription. At 20 minutes post infection, most of the mRNAs in the infected cell are viral mRNAs. The proteins translated from the early stage of transcription are involved in initiating viral DNA replication, occurring 60–90 minutes post infection. The second phase of proteins are translated in the cytoplasm and the assembly of virus capsids begins about 2–3 hours post infection. Mature virions are formed with the addition of newly replicated viral DNA from the host nucleus, likely facilitated by a virus encoded DNA packaging ATPase. About 5–6 hours following PBCV-1 infection, the cytoplasm is filled with virions and lysis occurs at 6–8 hours post infection releasing roughly 1000 particles per cell.[32][35]
Prymnesiovirus
[edit]The genus Prymnesiovirus currently contains only one species, known as Chrysochromulina brevifilum virus PW1 (CbV-PW1). CbV-PW1 infects two species of marine phytoplankton, Chrysochromulina brevifilum and C. strobilus, belonging to the genus Chrysochromulina.[36][37] According to the AlgaeBase database, there are currently 63 marine and freshwater species names in the genus, of which 48 are recognized as taxonomically acceptable names.[38] Chrysochromulina is a particularly important genus as it can comprise more than 50% of the photosynthetic nanoplanktonic cells in the ocean.[36]
Little is known about the life cycle of the virus infecting these flagellate-containing planktonic species, Chrysochromulina brevifilum and C. strobilus. Suttle and Chan (1995) were the first to isolate viruses which infect Prymnesiophytes or haptophytes. In this study, ultrathin sections of viruses within Chyrsochromulina brevifilum were prepared and viewed using transmission electron microscopy.[36] Electron micrographs in the early stage of infection suggest that virus replication occurs in the cytoplasm within a viroplasm. A viroplasm is a localized area in the cytoplasm, or around the nucleus of the cell which serves as a 'viral replication factory'. The viroplasm contains components such as virus genetic material, host proteins and ribosomes necessary for replication. Virosomes are often surrounded by a membrane; the membrane surrounding the virosome contained in the infected cells in the study was found to consist of a fibrillar matrix.[36] Virions are released from infected cells following disruption of the organelles and lysis of the host cell membrane. Suttle and Chan (1995) counted more than 320 viruses in an ultrathin section of an infection cell.[36] Estimates for burst sizes range from 320 to 600 viruses per cell.[39]
Prasinovirus
[edit]Members of the genus Prasinovirus infect small unicellular green algae in the order Mamiellales, commonly found in coastal marine waters.[40] A species of the genus Prasinovirus is Micromonas pusilla virus SP1 (MpV-SP1),[41] which was isolated from a water sample collected off of San Diego.[42] The prasinovirus MpV-SP1 infects Micromonas pusilla which is a dominant photosynthetic marine picoeukaryote.[43] and which infects Micromonas pusilla (UTEX 991, Plymouth 27). Common hosts of prasinoviruses include members from the genera Ostreococcus and Micromonas. Three potential species of Ostreococcus have been identified and differ based on their light requirements.[44] One of the most widely studied prasinoviruses, strain OtV5 whose genome is fully sequenced infects Ostreococcus tauri, the smallest free-living eukaryotes currently known.[45]
Prasinoviruses employ a nucleo-cytoplasmic replication strategy where virions adhere to the host-cell surface, followed by injection of DNA into the host cytoplasm.[45] Researchers found that 'empty' OtV5 viruses, or viruses with only the capsid attached to the host membrane, were rarely seen at any stage of the infection, suggesting that virions detach from the host membrane after injection of their DNA. The authors also found that a high proportion of viruses did not attach to cells after inoculation and suggest that viral attachment may be a limiting step in the infection. The viral DNA is then replicated inside the nucleus by the host cell's machinery. Virus particles are assembled in the cytoplasm, usually occupying a space near the inner face of the nucleus. Due to the extremely small size of the algae cells, the average burst size was found to be 25 virus particles per cell.[45]
Viral production without cell lysis has recently been observed in O. tauri cells. Thomas et al. (2011) found that in resistant host cells, the viral genome was replicated and viruses were released via a budding mechanism.[46] This low rate of viral release through budding allows for prolonged survivability of the host and virus progeny, resulting in a stable co-existence.[47]
Encoded proteins
[edit]Ectocarpus siliculosus virus (EsV-1), belonging to the genus Phaeovirus, and Paramecium bursaria chlorella virus (PBCV-1), belonging to the genus Chlorovirus, are two well-studied viruses, whose genomes have been found to encode many proteins. These proteins function in virus stability, DNA synthesis, transcription, and other important interactions with the host.[citation needed]
Enzymes for glycosylation
[edit]PBCV-1 has a 54-kDa glycosylated major capsid protein, which comprises about 40% of total viral protein.[12] Unlike most of the viral structural proteins which are glycosylated in the endoplasmic reticulum (ER) and Golgi apparatus by host-encoded glycosyltransferases,[48] PBCV-1 glycosylates its major capsid protein independently by encoding most of the enzymes necessary for constructing the complex oligosaccharides, which then attach to the major capsid protein of PBCV-1 to form the glycoprotein. Therefore, the glycosylation of the major capsid protein of PBCV-1 happens independently of the ER and Golgi apparatus in host cells.[49]
Ion channel proteins
[edit]The first known viral protein that functions as a potassium-selective ion channel was found in PBCV-1.[50] The protein (called Kcv) consists of 94 amino acids and is encoded from a small open reading frame (ORF) (ORF A250R) in PBCV-1, which can produce potassium-selective and voltage-sensitive conductance in Xenopus oocytes.[50] The supposed PBCV-1 protein has a short cytoplasmic N-terminus (12 amino acids) containing one consensus protein kinase C site and it has 2 transmembrane domains. The different amino acid sequences and lack of COOH-terminal cytoplasmic tail make the Kcv protein different from other potassium channels.[50][29]
EsV-1 encodes a 124 codon ORF that has significant amino acid similarity to PBCV-1 Kcv (41% amino acid identity).[29] However, the EsV-1 protein has a longer N-terminus (35 amino acids) containing two consensus protein kinase C sites and it has three transmembrane domains.[29] It is unknown whether the EsV-1 protein can form a functional channel in heterologous cells. The EsV-1 genome also encodes several proteins with hydrophobic amino acid rich regions that resemble helical transmembrane domains. Among these proteins, the input domain of the supposed hybrid His-kinase 186 and the ORF 188 resemble ion channel proteins.[45]
DNA replication-associated proteins
[edit]Both EsV-1 and PBCV-1 encode DNA polymerase which belong to the DNA polymerase-δ family, and they all contain a proof-reading 3'-5' exonuclease domain.[51] Additionally, both PBCV-1 and EsV-1 encode a sliding clamp processivity factor protein (PCNA), which interacts with proteins involved in DNA replication as well as proteins involved in DNA repair and postreplicative processing (e.g. DNA methylases and DNA transposases).[52]
Heteropentameric replication factor C (RFC) is a complex which is responsible for the ATP-dependent loading of PCNA onto DNA;[53][54] EsV-1 encodes five proteins which can form a RFC complex. PBCV-1 encodes a single protein which resembles the found in the Archae RFC complex.[45] PBCV-1 also encodes other proteins involved in DNA replication including an ATP-dependent DNA ligase,[55] a type II DNA topoisomerase, and RNase H.[29] Although both EsV-1 and PBCV-1 possess genes for essential elements of the eukaryotic replication system, neither have complete replicative genes, since they all lack genes for primase.[12][29]
Transcription-associated proteins
[edit]Neither EsV-1 nor PBCV-1 encode a complete RNA polymerase, but they produce several transcription factor-like proteins to assist the host transcription system.[citation needed]
EsV-1 encodes two small polypeptides (ORF 193 and ORF 196) for transcriptional regulation; the proteins resemble the α/β/α domain of TFIID-18 subunit.[45] The TFIID complex is necessary for transcription of eukaryotes, as it binds to the TATA box in the core promoter of the gene to initiate the assembly of RNA polymerase. Besides, polypeptides resemble to the SET, BTB/POZ (i.e. Broad Complex, Tramtrack, and Bric-a-brac/poxvirus and zinc finger) (ORF 40), and BAF60b (ORF 129) domains are also encoded by ESV-1 to regulate chromatin remodeling and transcription repression.[45][12][56]
Four transcription factor-like proteins have been found in PBSV-1, including TFIIB (A107L), TFIID (A552R), TFIIS (A125L), and a VLTF-2 type transcription factor (A482R).[29] In addition, PBCV-1 also encodes two enzymes involved in forming a mRNA cap structure, an RNA triphosphatase[57] and a mRNA guanylyltransferase.[58] The PBCV-1 enzymes are more closely related to yeast enzymes than to poxvirus multifunctional RNA capping enzymes according to its size, amino-acid sequence, and biochemical properties.[59][58] PBCV-1 also encodes RNase III, which is involved in virus mRNAs processing.[29]
Nucleotide metabolism-associated proteins
[edit]To supply deoxynucleotides for viral production in the low proliferating host cells, large DNA viruses possess genes to encode deoxynucleotide synthesis enzymes themselves.[29] Thirteen nucleotide metabolic enzymes have been found in PBCV-1, two of which include dUTP pyrophosphatase and dCMP deaminase, which can produce dUMP (i.e. the substrate for thymidylate synthetase).[60] In comparison, EsV-1 only encodes an ATPase (ORF 26) as well as both subunits of ribonucleotide reductase (ORF 128 and 180), which is a key enzyme in deoxynucleotide synthesis.[45]
Other enzymes
[edit]Other enzymes such as methyltransferases, DNA restriction endonucleases, and integrase were also found in PBCV-1.[12][29] PBCV-1 also encodes a 187-amino-acid protein that resembles the Cu-Zn SOD with all of the conserved amino acid residues for binding copper and zinc, which can decompose the rapid accumulated superoxide in host cells during infection, thereby benefiting virus replication.[61]
Ecological implications
[edit]Raphidovirus
[edit]
Heterosigma akashiwo forms dense, harmful blooms in temperate and subarctic waters, occurring at densities up to 5 ×106 cells/ml.[62] These algal blooms can be extremely harmful to aquatic life, causing mortality in wild and cultured fish, such as salmon, yellowtail and sea bream.[5] The severity and duration of these blooms varies from year, and damage to aquaculture by H.akashiwo has been increasing. In 1989, a noxious algal bloom off the coast of New Zealand resulted in the loss of seventeen million New Zealand dollars worth of Chinook salmon. In 1995 and 1997 in Japanese coastal waters in Kagoshimo Bay, 1,090 million and 327 million Yen worth of fish were killed, respectively.[5]
The HaV virus, infecting H. akashiwo has been shown to be a factor in bloom termination. Suttle et al. (1990) suggested that viral infection of algae could have a role in regulating population densities of phytoplankton communities, thus having significant roles in their dynamics in the oceans.[63] Earlier studies, such as the study by Nagasaki et al. (1993), explored the dynamics between HaV and H. akashiwo. Algal samples were obtained in the middle or final stages of a red tide in Hiroshima Bay, Japan. Using transmission electron microscopy, Nagaski et al. identified the HaV virus in and around the nuclear area of H. akashiwo cells.[63] Further support for the role of the HaV virus in bloom termination was provided by a study conducted by Nagaski et al. (1994). Nagaski et al. (1994) found that proportion of virus-containing cells increased quickly before termination of the red tide; no virus-containing cells were detected three days before termination of the red tide and the sample collected on the last day revealed a high frequency (11.5%) of virus-containing cells.[64]
Further studies by Tarutani et al. (2000) also found an association between a decrease in cell density of H. akashiwo with an increase in the abundance of HaV. The researchers found that HaV not only plays in important role in controlling biomass, but also influences the clonal composition or characteristics of H. akashiwo cells. The researchers found that most isolates following bloom termination were resistant to HaV clonal isolates, while during bloom formation resistant cells were a minor component. The authors suggest that viral infection, during the bloom termination period influences the properties of dominant cells in H. akashiwo populations.[65] Selective pressure exerted by the viruses in the later stage of infection may promote genetic diversity, allowing the H. akashiwo population to thrive after bloom termination.[citation needed]
As mentioned, H. akashiwo blooms are detrimental to fish populations in temperate and subarctic waters, and continue to pose serious threats for aquaculture. Nagasaki et al. (1999) examined the growth characteristics of HaV and suggested that HaV could be used as a microbial agent against H. akashiwo red tides. The advantages of using HaV is that it specifically infects H. akashiwo even when other microorganisms are present. Additionally, it has a high growth rate and can be produced at a low cost. Using HaV as a microbial agent is a promising solution for eliminating red tides to protect fisheries and marine life, but as the authors concluded, the effects of various HaV clones on H. akashiwo populations should be explored in greater detail before the virus is used for wide-scale applications.[5]
Coccolithovirus
[edit]
The coccolithovirus (EhV) infects the coccolithophore Emiliania huxleyi (E. huxleyi). Coccolithophores are marine haptophytes which are surrounded by microscopic plates made of calcium carbonate.[66] They live in the upper layers of the world's oceans and represent the third most abundant group of phytoplankton, containing about 300 species.[67] E. huxleyi is recognized as the most prominent and ecologically important of the coccolithophores. E. huxleyi has a global distribution from the tropics to subarctic waters and occasionally forms dense blooms which can cover 100,000s of square kilometers.[67] These trillions of coccolithophores produced, then die and sink to the bottom of oceans, contributing to sediment formation, and are the biggest producers of calcite in the oceans.[66] Thus, coccoliths have significant roles in global carbon fixation and the carbon cycle as well as sulfur cycling.[67] Over time, coccolithophores have shaped geological features of our planet. For example, the White Cliffs of Dover are formed from white chalk, or calcium carbonate produced by coccolithophores over millions of years.[citation needed]
Coccolithophore blooms are typically not harmful to marine life in the ocean. As these organisms thrive in nutrient-poor conditions, the coccolithophores offer a source of nutrition for small fish and zooplankton.[66] E. huxylei viruses (EhVs) have been shown to be linked to the termination of these blooms. The termination stage of the bloom is indicated by a color change in the water. When large amounts of coccoliths (carbonate shell surrounding E. huxylei) are shed from E. huxylei cells from cell death or lysis, the water turns white or turquoise. In areas of dense bloom termination, the white color is reflective and can be seen in satellite imagery.[67] Wilson et al. (2002) used analytical flow cytometry to measure the abundance of viruses at different locations in and around the bloom area. The researchers found that the concentrations of viruses were higher inside the 'high reflectance area', suggesting that virus-induced lysis of E. huxleyi cells resulted in coccolith detachment.[68] Other studies by Martinez et al. (2007) and Bratbak et al. (1993) found higher concentrations of EhV viruses as the E. huxleyi bloom declined, indicating that lytic viral infection was the main cause of bloom termination.[69][70] EhV viruses therefore have important roles in regulating biomass production in marine environments and ecological succession. This regulation of coccolithophore populations by EhV viruses therefore has significant effects on biogeochemical cycles, particularly the carbon cycle.[citation needed]
Phaeovirus
[edit]
One of the best-studied phaeoviruses, EsV-1, infects the small, filamentous brown algae E. siliculosus, which has a cosmopolitan distribution (found in most of the world's oceans).[29] The Ectocarpales are closely related to the brown algal group, the Laminariales, which are the most economically important group of brown algae, having a wide range of applications in the cosmetics and food industry.[71]
Muller et al. (1990) were one of the first to explore the causes of gametangium defects in E. siliculosus originating from New Zealand. The researchers identified reproductive cells of E. siliculosus filled with hexagonal particles which were then released into culture medium when the cells burst. Following release of these particles, sporophytes became infected, shown by pathological symptoms, suggesting that the particles are viruses.[72] Such studies allowed for the evaluation of infection potential of E. siliculosus viruses. Using PCR amplification of a viral gene fragment, Muller et al. (2005) monitored levels of pathogen infection in Ectocarpus samples from the Gran Canaria Island, North Atlantic and southern Chile. The researchers found high levels of pathogen prevalence; 40–100% of Ectocarpus specimens contained viral DNA.[73] Similar estimates have been given by Sengco et al. (1996) who estimated that at least 50% of Ectocarpus plants in the world contain viral DNA.[74] This high frequency of viral infection among globally distributed Ectocarpus plants has ecological implications. Viral infection by EsV-1 in E. siliculosus plants, as mentioned, limits reproductive success of infected plants. Thus, the EsV-1 virus plays a key role in regulating populations of E. siliculosus, having further effects on local ecosystem dynamics.[citation needed]
Chlorovirus
[edit]
Members of the genus Chlorovirus are found in freshwater sources around the world and infect the green algae symbionts zoochlorellae. There is a lack of information about the role chloroviruses play in freshwater ecology.[75] Despite this, chloroviruses are found in native waters at 1–100 plaque-forming units (PFU)/ml and measurements as high as 100,000 PFU/ml of native water have been obtained.[8] A plaque-forming unit is the number of particles capable of forming visible structures within a cell culture, known as plaques.[citation needed] Abundances of chloroviruses vary with season, with the highest abundances occurring in the spring.[8] Chloroviruses, such as PBCV-1, play a role in regulating host populations of zoochlorella. As mentioned previously, infection of zoochlorella occurs only when the symbiotic relationship with its host is disrupted. Infection of the algae during this stage of host/algae independence will prevent the host and algae relationship from being restored, thus decreasing the survivability of the endosymbiotic hosts of the zoochlorellae, such as Paramecium bursaria. Thus, chloroviruses play in important role in freshwater ecosystems by not only regulating populations of their host, zoochlorellae, but also regulating, to an extent, populations of zoochlorellae hosts as well. Chloroviruses and viruses in general cause death and lysis of their hosts, releasing dissolved organic carbon, nitrogen and phosphorus into the water. These nutrients can then be taken up by bacteria, thus contributing to the microbial loop. Liberation of dissolved organic materials allows for bacterial growth, and bacteria are an important source of food for organisms in higher trophic levels. Consequently, chloroviruses have significant effects on carbon and nutrient flows, influencing freshwater ecosystem dynamics.[6]
Prymnesiovirus
[edit]Prymnesiovirus, CbV-PW1, as mentioned infects the algal genus Chyrsochromulina. Chyrsochromulina, found in global fresh and marine waters, occasionally forms dense blooms which can produce harmful toxins, having negative effects on fisheries.[36] A particularly toxic species called C. polylepis has caused enormous damage to commercial fisheries in Scandinavia. In 1988, this bloom caused a loss of 500 tons of caged fish, worth 5 million US.[76] Given that Chyrsochromulina is a widespread species, and is of significant ecological importance, viral infection and lysis of genus members is likely to have significant impacts on biogeochemical cycles, such as nutrient recycling in aquatic environments. Suttle and Chan suggest that the presence of viruses should have a strong regulatory effect on Chyrsochromulina populations, thus preventing bloom formation or enabling bloom termination, explaining why persistent blooms are an unusual phenomenon in nature.[36]
Prasinovirus
[edit]A commonly studied prasinovirus, OtV5, as mentioned, infects the smallest currently known eukaryote, Ostreococcus tauri. O. tauri is about 0.8 micrometers in diameter and is within the picosize fraction (0.2–2 micrometers). Picoeukaryotes, such as Ostreococcus tauri are widely distributed and contribute significantly to microbial biomass and total primary productivity. In oligotrophic environments, marine picophytoplankton account for up to 90% of the autotrophic biomass and thus are an important food source for nanoplanktonic and phagotrophic protists.[77] As picoeukaryotes serve as the base for marine microbial food webs, they are intrinsic to the survival of higher trophic levels. Ostreococcus tauri has a rapid growth rate and dense blooms have been observed off the coasts of Long Island and California.[77] Samples collected from Long Island bay were found to contain many virus-like particles, a likely cause for the decline of the bloom.[78] Despite the large abundances of picoeukaryotes, these unicellular organisms are outnumbered by viruses by about ten to one.[79] Viruses such as OtV5, play important roles in regulating phytoplankton populations, and through lysis of cells contribute to the recycling of nutrients back towards other microorganisms, otherwise known as the viral shunt.[80]
As mentioned, the prasinovirus MpV-SP1 infects Micromonas pusilla which is a major component of the picophytoplankton of the world's oceans. M. pusilla lives from tropical to polar marine ecosystems.[81] Cottrell & Suttle (1995) found that 2–10% of the M. pusilla population in an inshore environment was lysed per day, with an average of 4.4%.[43] Higher estimates have been given by Evans et al. (2003), who suggest that M. pusilla viruses can lyse up to 25% of the Micromonas population per day.[82] This suggests that viruses are responsible for a moderate amount of mortality in M. pusilla populations.[43] On a larger scale, viral infection of M. pusilla is responsible for nutrient and energy recycling in aquatic food webs, which is yet to be quantified.[citation needed]
Pathology
[edit]Until recently phycodnaviruses were believed to infect algal species exclusively. Recently, DNA homologous to Chlorovirus Acanthocystis turfacea virus 1 (ATCV-1) were isolated from human nasopharyngeal mucosal surfaces. The presence of ATCV-1 in the human microbiome was associated with diminished performance on cognitive assessments. Inoculation of ATCV-1 in experimental animals was associated with decreased performance in memory and sensory-motor gating, as well as altered expression of genes in the hippocampus related to synaptic plasticity, learning, memory formation, and the viral immune response.[3]
References
[edit]- ^ "Viral Zone". ExPASy. Retrieved 15 June 2015.
- ^ a b ICTV. "Virus Taxonomy: 2014 Release". Retrieved 15 June 2015.
- ^ a b Yolken, RH; et al. (2014). "Chlorovirus ATCV-1 is part of the human oropharyngeal virome and is associated with changes in cognitive functions in humans and mice". Proc Natl Acad Sci U S A. 111 (45): 16106–16111. Bibcode:2014PNAS..11116106Y. doi:10.1073/pnas.1418895111. PMC 4234575. PMID 25349393.
- ^ a b c d e Tomaru, Yuji; Shirai, Yoko; Nagasaki, Keizo (1 August 2008). "Ecology, physiology and genetics of a phycodnavirus infecting the noxious bloom-forming raphidophyte Heterosigma akashiwo". Fisheries Science. 74 (4): 701–711. Bibcode:2008FisSc..74..701T. doi:10.1111/j.1444-2906.2008.01580.x. S2CID 23152411.
- ^ a b c d e Nagasaki, Keizo; Tarutani, Kenji; Yamaguchi, Mineo (1 March 1999). "Growth Characteristics of Heterosigma akashiwo Virus and Its Possible Use as a Microbiological Agent for Red Tide Control". Applied and Environmental Microbiology. 65 (3): 898–902. Bibcode:1999ApEnM..65..898N. doi:10.1128/AEM.65.3.898-902.1999. PMC 91120. PMID 10049839.
- ^ a b Sigee, David (27 September 2005). Freshwater Microbiology: Biodiversity and Dynamic Interactions of Microorganisms in the Aquatic Environment. John Wiley & Sons. ISBN 9780470026472.
- ^ Anonymous (2012) Virus Taxonomy: IXth report of the international committee on taxonomy of viruses. Amsterdam: Academic Press, p. 261.
- ^ a b c d e f g Dunigan, David D.; Fitzgerald, Lisa A.; Van Etten, James L. (2006). "Phycodnaviruses: A peek at genetic diversity". Virus Research. 117 (1): 119–132. doi:10.1016/j.virusres.2006.01.024. PMID 16516998. S2CID 11532538.
- ^ Chen, Feng; Suttle, Curtis A (1996). "Evolutionary relationships among large double-stranded DNA viruses that infect microalgae and other organisms as inferred from DNA polymerase genes". Virology. 219 (1): 170–178. doi:10.1006/viro.1996.0234. PMID 8623526.
- ^ Baker, Timothy S.; Yan, Xiaodong; Olson, Norman H.; Etten, James L. Van; Bergoin, Max; Rossmann, Michael G. (2000). "Nature Citation". Nature Structural Biology. 7 (2): 101–103. doi:10.1038/72360. PMC 4167659. PMID 10655609.
- ^ Yan, X.; Chipman, P. R.; Castberg, T.; Bratbak, G.; Baker, T. S. (2005). "The Marine Algal Virus PpV01 Has an Icosahedral Capsid with T=219 Quasisymmetry". Journal of Virology. 79 (14): 9236–9243. doi:10.1128/JVI.79.14.9236-9243.2005. PMC 1168743. PMID 15994818.
- ^ a b c d e f g Mackinder, Luke C. M.; Worthy, Charlotte A.; Biggi, Gaia; Hall, Matthew; Ryan, Keith P.; Varsani, Arvind; Harper, Glenn M.; Wilson, William H.; Brownlee, Colin (1 January 2009). "A unicellular algal virus, Emiliania huxleyi virus 86, exploits an animal-like infection strategy". Journal of General Virology. 90 (9): 2306–2316. doi:10.1099/vir.0.011635-0. PMID 19474246.
- ^ Viruses, International Committee on Taxonomy of; King, Andrew M.Q. (2012). "Phycodnaviridae". Virus Taxonomy. pp. 249–262. doi:10.1016/B978-0-12-384684-6.00024-0. ISBN 9780123846846.
- ^ Li, Yu; Lu, Zhiqiang; Sun, Liangwu; Ropp, Susan; Kutish, Gerald F.; Rock, Daniel L.; Van Etten, James L. (1997). "Analysis of 74 kb of DNA Located at the Right End of the 330-kb Chlorella Virus PBCV-1 Genome". Virology. 237 (2): 360–377. doi:10.1006/viro.1997.8805. PMID 9356347.
- ^ a b Delaroque, Nicolas; Müller, Dieter Gerhard; Bothe, Gordana; Pohl, Thomas; Knippers, Rolf; Boland, Wilhelm (2001). "The Complete DNA Sequence of the Ectocarpus siliculosus Virus EsV-1 Genome". Virology. 287 (1): 112–132. doi:10.1006/viro.2001.1028. PMID 11504547.
- ^ a b c Wilson, W. H.; Schroeder, D. C.; Allen, M. J.; Holden, M. T.; Parkhill, J.; Barrell, B. G.; Churcher, C.; Hamlin, N.; Mungall, K.; Norbertczak, H.; Quail, M. A.; Price, C.; Rabbinowitsch, E.; Walker, D.; Craigon, M.; Roy, D.; Ghazal, P. (2005). "Complete Genome Sequence and Lytic Phase Transcription Profile of a Coccolithovirus". Science. 309 (5737): 1090–1092. Bibcode:2005Sci...309.1090W. doi:10.1126/science.1113109. PMID 16099989. S2CID 6710771.
- ^ Finke, Jan; Winget, Danielle; Chan, Amy; Suttle, Curtis (2017). "Variation in the Genetic Repertoire of Viruses Infecting Micromonas pusilla Reflects Horizontal Gene Transfer and Links to Their Environmental Distribution". Viruses. 9 (5): 116. doi:10.3390/v9050116. PMC 5454428. PMID 28534829.
- ^ Allen, M. J.; Schroeder, D. C.; Wilson, W. H. (1 March 2006). "Preliminary characterisation of repeat families in the genome of EhV-86, a giant algal virus that infects the marine microalga Emiliania huxleyi". Archives of Virology. 151 (3): 525–535. doi:10.1007/s00705-005-0647-1. PMID 16195784. S2CID 10421212.
- ^ Allen, Michael J.; Schroeder, Declan C.; Donkin, Andrew; Crawfurd, Katharine J.; Wilson, William H. (2006). "Genome comparison of two Coccolithoviruses". Virology Journal. 3: 15. doi:10.1186/1743-422X-3-15. PMC 1440845. PMID 16553948.
- ^ Kumar, Sudhir; Stecher, Glen; Tamura, Koichiro (2016). "MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets". Molecular Biology and Evolution. 33 (7): 1870–1874. doi:10.1093/molbev/msw054. PMC 8210823. PMID 27004904.
- ^ a b Maruyama, Fumito; Ueki, Shoko (2016). "Evolution and Phylogeny of Large DNA Viruses, Mimiviridae and Phycodnaviridae Including Newly Characterized Heterosigma akashiwo Virus". Frontiers in Microbiology. 7: 1942. doi:10.3389/fmicb.2016.01942. PMC 5127864. PMID 27965659.
- ^ Fischer, M. G.; Allen, M. J.; Wilson, W. H.; Suttle, C. A. (2010). "Giant virus with a remarkable complement of genes infects marine zooplankton". Proceedings of the National Academy of Sciences. 107 (45): 19508–19513. Bibcode:2010PNAS..10719508F. doi:10.1073/pnas.1007615107. PMC 2984142. PMID 20974979.
- ^ Yau, S.; Lauro, F. M.; Demaere, M. Z.; Brown, M. V.; Thomas, T.; Raftery, M. J.; Andrews-Pfannkoch, C.; Lewis, M.; Hoffman, J. M.; Gibson, J. A.; Cavicchioli, R. (2011). "Virophage control of antarctic algal host-virus dynamics". Proceedings of the National Academy of Sciences. 108 (15): 6163–6168. Bibcode:2011PNAS..108.6163Y. doi:10.1073/pnas.1018221108. PMC 3076838. PMID 21444812.
- ^ Hull, Roger (2014). "Plant Viruses and Their Classification". Plant Virology. pp. 15–68. doi:10.1016/B978-0-12-384871-0.00002-9. ISBN 9780123848710.
- ^ Tai, Vera; Lawrence, Janice E; Lang, Andrew S; Chan, Amy M; Culley, Alexander I; Suttle, Curtis A (2003). "Characterization of HaRNAV, a single-stranded RNA virus causing lysis of Heterosigma akashiwo (Raphidophyceae)". Journal of Phycology. 39 (2): 343–352. doi:10.1046/j.1529-8817.2003.01162.x. S2CID 86464306.
- ^ Lawrence, Janice E; Chan, Amy M; Suttle, Curtis A (2001). "A novel virus (HaNIV) causes lysis of the toxic bloom-forming alga Heterosigma akashiwo (Raphidophyceae)". Journal of Phycology. 37 (2): 216–222. Bibcode:2001JPcgy..37..216L. doi:10.1046/j.1529-8817.2001.037002216.x. S2CID 86563435.
- ^ Vardi, A.; Van Mooy, B. A. S.; Fredricks, H. F.; Popendorf, K. J.; Ossolinski, J. E.; Haramaty, L.; Bidle, K. D. (2009). "Viral Glycosphingolipids Induce Lytic Infection and Cell Death in Marine Phytoplankton". Science. 326 (5954): 861–865. Bibcode:2009Sci...326..861V. doi:10.1126/science.1177322. PMID 19892986. S2CID 40102053.
- ^ a b Stevens, Kim; Weynberg, Karen; Bellas, Christopher; Brown, Sonja; Brownlee, Colin; Brown, Murray T.; Schroeder, Declan C. (2014). "A Novel Evolutionary Strategy Revealed in the Phaeoviruses". PLOS ONE. 9 (1): e86040. Bibcode:2014PLoSO...986040S. doi:10.1371/journal.pone.0086040. PMC 3897601. PMID 24465858.
- ^ a b c d e f g h i j k l Klein, M.; Lanka, S. T.; Knippers, R.; Müller, D. G. (10 January 1995). "Coat protein of the Ectocarpus siliculosus virus". Virology. 206 (1): 520–526. doi:10.1016/s0042-6822(95)80068-9. PMID 7831806.
- ^ a b Charrier, Bénédicte; Coelho, Susana M.; Le Bail, Aude; Tonon, Thierry; Michel, Gurvan; Potin, Philippe; Kloareg, Bernard; Boyen, Catherine; Peters, Akira F.; Cock, J. Mark (2007). "Development and physiology of the brown alga Ectocarpus siliculosus: Two centuries of research" (PDF). New Phytologist. 177 (2): 319–332. doi:10.1111/j.1469-8137.2007.02304.x. PMID 18181960. S2CID 43432914.
- ^ a b Müller, DG (1991). "Marine virioplankton produced by infected Ectocarpus siliculosus (Phaeophyceae)". Marine Ecology Progress Series. 76: 101–102. Bibcode:1991MEPS...76..101M. doi:10.3354/meps076101.
- ^ a b Sigee, David (27 September 2005). Freshwater Microbiology: Biodiversity and Dynamic Interactions of Microorganisms in the Aquatic Environment. John Wiley & Sons. ISBN 9780470026472.
- ^ Etten, James L. Van; Dunigan, David D. (18 August 2016). "Giant Chloroviruses: Five Easy Questions". PLOS Pathogens. 12 (8): e1005751. doi:10.1371/journal.ppat.1005751. PMC 4990331. PMID 27536965.
- ^ Delong, John P.; Al-Ameeli, Zeina; Duncan, Garry; Van Etten, James L.; Dunigan, David D. (2016). "Predators catalyze an increase in chloroviruses by foraging on the symbiotic hosts of zoochlorellae". Proceedings of the National Academy of Sciences. 113 (48): 13780–13784. Bibcode:2016PNAS..11313780D. doi:10.1073/pnas.1613843113. PMC 5137705. PMID 27821770.
- ^ Van Etten, James L.; Dunigan, David D. (2012). "Chloroviruses: Not your everyday plant virus". Trends in Plant Science. 17 (1): 1–8. doi:10.1016/j.tplants.2011.10.005. PMC 3259250. PMID 22100667.
- ^ a b c d e f g Suttle, CA; Chan, AM (1995). "Viruses infecting the marine Prymnesiophyte Chrysochromulina spp.: Isolation, preliminary characterization and natural abundance". Marine Ecology Progress Series. 118: 275–282. Bibcode:1995MEPS..118..275S. doi:10.3354/meps118275.
- ^ Suttle, Curtis A.; Chan, Amy M. (2002). "Prymnesiovirus". The Springer Index of Viruses. pp. 741–743. doi:10.1007/3-540-31042-8_128. ISBN 978-3-540-67167-1.
- ^ "Chrysochromulina Lackey, 1939 :: Algaebase". www.algaebase.org. Retrieved 28 February 2017.
- ^ King, Andrew M. Q. (1 January 2012). Virus Taxonomy: Classification and Nomenclature of Viruses : Ninth Report of the International Committee on Taxonomy of Viruses. Elsevier. ISBN 9780123846846.
- ^ Clerissi, Camille; Desdevises, Yves; Grimsley, Nigel (2012). "Prasinoviruses of the Marine Green Alga Ostreococcus tauri Are Mainly Species Specific". Journal of Virology. 86 (8): 4611–4619. doi:10.1128/JVI.07221-11. PMC 3318615. PMID 22318150.
- ^ "ICTV Taxonomy history: Micromonas pusilla virus SP1". ICTV. 8 March 1998. Retrieved 25 June 2017.
- ^ Cottrell, MT; Suttle, CA (1991). "Wide-spread occurrence and clonal variation in viruses which cause lysis of a cosmopolitan, eukaryotic marine phytoplankter Micromonas pusilla". Marine Ecology Progress Series. 78: 1–9. Bibcode:1991MEPS...78....1C. doi:10.3354/meps078001.
- ^ a b c Cottrell, Matthew T.; Suttle, Curtis A. (1995). "Dynamics of lytic virus infecting the photosynthetic marine picoflagellate Micromonas pusilla". Limnology and Oceanography. 40 (4): 730–739. Bibcode:1995LimOc..40..730C. doi:10.4319/lo.1995.40.4.0730.
- ^ "HomeOstreococcus lucimarinus". genome.jgi.doe.gov. Retrieved 28 February 2017.
- ^ a b c d e f g h Derelle, Evelyne; Ferraz, Conchita; Escande, Marie-Line; Eychenié, Sophie; Cooke, Richard; Piganeau, Gwenaël; Desdevises, Yves; Bellec, Laure; Moreau, Hervé (28 May 2008). "Life-Cycle and Genome of OtV5, a Large DNA Virus of the Pelagic Marine Unicellular Green Alga Ostreococcus tauri". PLOS ONE. 3 (5): e2250. Bibcode:2008PLoSO...3.2250D. doi:10.1371/journal.pone.0002250. PMC 2386258. PMID 18509524.
- ^ Thomas, Rozenn; Grimsley, Nigel; Escande, Marie-Line; Subirana, Lucie; Derelle, Evelyne; Moreau, Hervé (2011). "Acquisition and maintenance of resistance to viruses in eukaryotic phytoplankton populations". Environmental Microbiology. 13 (6): 1412–1420. Bibcode:2011EnvMi..13.1412T. doi:10.1111/j.1462-2920.2011.02441.x. PMID 21392198.
- ^ Sime-Ngando, TéLesphore (2014). "Environmental bacteriophages: Viruses of microbes in aquatic ecosystems". Frontiers in Microbiology. 5: 355. doi:10.3389/fmicb.2014.00355. PMC 4109441. PMID 25104950.
- ^ Sigvard Olofsson, John-Erik s. Hans (1998). "Host Cell Glycosylation of Viral Glycoproteins - a Battlefield for Host Defence and Viral Resistance". Scandinavian Journal of Infectious Diseases. 30 (5): 435–440. doi:10.1080/00365549850161386. PMID 10066039.
- ^ Markine-Goriaynoff, N.; Gillet, L.; Van Etten, J. L.; Korres, H.; Verma, N.; Vanderplasschen, A. (2004). "Glycosyltransferases encoded by viruses". Journal of General Virology. 85 (10): 2741–2754. doi:10.1099/vir.0.80320-0. PMID 15448335.
- ^ a b c Plugge, B.; Gazzarrini, S.; Nelson, M.; Cerana, R.; Van Etten, J. L.; Derst, C.; Difrancesco, D.; Moroni, A.; Thiel, G. (2000). "A Potassium Channel Protein Encoded by Chlorella Virus PBCV-1". Science. 287 (5458): 1641–1644. Bibcode:2000Sci...287.1641P. doi:10.1126/science.287.5458.1641. PMID 10698737. S2CID 17411728.
- ^ Hübscher, Ulrich; Nasheuer, Heinz-Peter; Syväoja, Juhani E. (2000). "Eukaryotic DNA polymerases, a growing family". Trends in Biochemical Sciences. 25 (3): 143–147. doi:10.1016/S0968-0004(99)01523-6. PMID 10694886.
- ^ Warbrick, E. (2000). "The puzzle of PCNA's many partners". BioEssays. 22 (11): 997–1006. doi:10.1002/1521-1878(200011)22:11<997::AID-BIES6>3.0.CO;2-#. PMID 11056476. S2CID 196597356.
- ^ Ellison, Viola; Stillman, Bruce (2001). "Opening of the Clamp". Cell. 106 (6): 655–660. doi:10.1016/S0092-8674(01)00498-6. PMID 11572772. S2CID 17575635.
- ^ Mossi, Romina; Hubscher, Ulrich (1998). "Clamping down on clamps and clamp loaders. The eukaryotic replication factor C". European Journal of Biochemistry. 254 (2): 209–216. doi:10.1046/j.1432-1327.1998.2540209.x. PMID 9660172.
- ^ Ho, C. K.; Van Etten, J. L.; Shuman, S. (1997). "Characterization of an ATP-dependent DNA ligase encoded by Chlorella virus PBCV-1". Journal of Virology. 71 (3): 1931–7. doi:10.1128/JVI.71.3.1931-1937.1997. PMC 191272. PMID 9032324.
- ^ Deweindt, C.; Albagli, O.; Bernardin, F.; Dhordain, P.; Quief, S.; Lantoine, D.; Kerckaert, J. P.; Leprince, D. (1995). "The LAZ3/BCL6 oncogene encodes a sequence-specific transcriptional inhibitor: A novel function for the BTB/POZ domain as an autonomous repressing domain". Cell Growth & Differentiation. 6 (12): 1495–503. PMID 9019154.
- ^ Ho, C. K.; Gong, C.; Shuman, S. (2001). "RNA Triphosphatase Component of the mRNA Capping Apparatus of Paramecium bursaria Chlorella Virus 1". Journal of Virology. 75 (4): 1744–1750. doi:10.1128/JVI.75.4.1744-1750.2001. PMC 114083. PMID 11160672.
- ^ a b Ho, C. K.; Van Etten, J. L.; Shuman, S. (1996). "Expression and characterization of an RNA capping enzyme encoded by Chlorella virus PBCV-1". Journal of Virology. 70 (10): 6658–64. doi:10.1128/JVI.70.10.6658-6664.1996. PMC 190707. PMID 8794301.
- ^ Shuman, S. (2001). "Structure, mechanism, and evolution of the mRNA capping apparatus". Progress in Nucleic Acid Research and Molecular Biology. 66: 1–40. doi:10.1016/s0079-6603(00)66025-7. ISBN 9780125400664. PMID 11051760.
- ^ Van Etten, James L.; Meints, Russel H. (1999). "Giant Viruses Infecting Algae". Annual Review of Microbiology. 53: 447–494. doi:10.1146/annurev.micro.53.1.447. PMID 10547698. S2CID 7346326.
- ^ Kang, M.; Duncan, G. A.; Kuszynski, C.; Oyler, G.; Zheng, J.; Becker, D. F.; Van Etten, J. L. (2014). "Chlorovirus PBCV-1 Encodes an Active Copper-Zinc Superoxide Dismutase". Journal of Virology. 88 (21): 12541–12550. doi:10.1128/JVI.02031-14. PMC 4248938. PMID 25142578.
- ^ Lawrence, Janice E.; Brussaard, Corina P. D.; Suttle, Curtis A. (1 March 2017). "Virus-Specific Responses of Heterosigma akashiwo to Infection". Applied and Environmental Microbiology. 72 (12): 7829–7834. Bibcode:2006ApEnM..72.7829L. doi:10.1128/AEM.01207-06. PMC 1694243. PMID 17041155.
- ^ a b Nagasaki, K.; Ando, M.; Imai, I.; Itakura, S.; Ishida, Y. (1994). "Virus-like particles in Heterosigma akashiwo (Raphidophyceae): a possible red tide disintegration mechanism". Marine Biology. 119 (2): 307–312. Bibcode:1994MarBi.119..307N. doi:10.1007/BF00349570. S2CID 85350528.
- ^ Nagasaki, Keizo; Ando, Masashi; Itakura, Shigeru; Imai, Ichiro; Ishida, Yuzaburo (1994). "Viral mortality in the final stage of Heterosigma akashiwo (Raphidophyceae) red tide". Journal of Plankton Research. 16 (11): 1595–1599. doi:10.1093/plankt/16.11.1595.
- ^ Tarutani, K.; Nagasaki, K.; Yamaguchi, M. (2000). "Viral Impacts on Total Abundance and Clonal Composition of the Harmful Bloom-Forming Phytoplankton Heterosigma akashiwo". Applied and Environmental Microbiology. 66 (11): 4916–4920. Bibcode:2000ApEnM..66.4916T. doi:10.1128/AEM.66.11.4916-4920.2000. PMC 92399. PMID 11055943.
- ^ a b c John, Weier (26 April 1999). "What is a Coccolithophore? Fact Sheet : Feature Articles". earthobservatory.nasa.gov. Retrieved 3 March 2017.
- ^ a b c d "Home—Emiliania huxleyi". genome.jgi.doe.gov. Retrieved 3 March 2017.
- ^ Wilson, William H.; Tarran, Glen A.; Schroeder, Declan; Cox, Michael; Oke, Joanne; Malin, Gillian (2002). "Isolation of viruses responsible for the demise of an Emiliania huxleyi bloom in the English Channel" (PDF). Journal of the Marine Biological Association of the UK. 82 (3): 369–377. Bibcode:2002JMBUK..82..369W. doi:10.1017/S002531540200560X. S2CID 85219045.
- ^ Martinez, J. M.; Schroeder, D. C.; Larsen, A.; Bratbak, G.; Wilson, W. H. (2007). "Molecular Dynamics of Emiliania huxleyi and Cooccurring Viruses during Two Separate Mesocosm Studies". Applied and Environmental Microbiology. 73 (2): 554–562. Bibcode:2007ApEnM..73..554M. doi:10.1128/AEM.00864-06. PMC 1796978. PMID 17098923.
- ^ Bratbak, G.; Egge, JK; Heldal, M. (1993). "Viral mortality of the marine alga Emiliania huxleyi (Haptophyceae) and termination of algal blooms". Marine Ecology Progress Series. 93: 39–48. Bibcode:1993MEPS...93...39B. doi:10.3354/meps093039.
- ^ "Ectocarpus siliculosus a genetic and genomic model organism for the brown algae". François Jacob Institute of biology. 21 June 2018.
- ^ Müller, D. G.; Kawai, H.; Stache, B.; Lanka, S. (1990). "A Virus Infection in the Marine Brown Alga Ectocarpus siliculosus(Phaeophyceae)". Botanica Acta. 103 (1): 72–82. doi:10.1111/j.1438-8677.1990.tb00129.x.
- ^ Müller, D. G.; Westermeier, R.; Morales, J.; Reina, G. Garcia; Del Campo, E.; Correa, J. A.; Rometscha, E. (2000). "Massive Prevalence of Viral DNA in Ectocarpus (Phaeophyceae, Ectocarpales) from Two Habitats in the North Atlantic and South Pacific". Botanica Marina. 43 (2). doi:10.1515/BOT.2000.016. hdl:10553/73463. S2CID 84166193.
- ^ Sengco, M. R.; Bräutigam, M.; Kapp, M.; Müller, D. G. (1 February 1996). "Detection of virus DNA in Ectocarpus siliculosus and E. fasciculatus (Phaeophyceae) from various geographic areas". European Journal of Phycology. 31 (1): 73–78. Bibcode:1996EJPhy..31...73S. doi:10.1080/09670269600651221.
- ^ Dunigan, David D.; Cerny, Ronald L.; Bauman, Andrew T.; Roach, Jared C.; Lane, Leslie C.; Agarkova, Irina V.; Wulser, Kurt; Yanai-Balser, Giane M.; Gurnon, James R.; Vitek, Jason C.; Kronschnabel, Bernard J.; Jeanniard, Adrien; Blanc, Guillaume; Upton, Chris; Duncan, Garry A.; McClung, O. William; Ma, Fangrui; Van Etten, James L. (2012). "Paramecium bursaria Chlorella Virus 1 Proteome Reveals Novel Architectural and Regulatory Features of a Giant Virus". Journal of Virology. 86 (16): 8821–8834. doi:10.1128/JVI.00907-12. PMC 3421733. PMID 22696644.
- ^ "Toxic Algal Bloom in Scandinavian Waters, May–June 1988 | Oceanography". Oceanography. 2 (1): 9–14. 1989. doi:10.5670/oceanog.1989.24. Retrieved 1 March 2017.
- ^ a b Derelle, E.; Ferraz, C.; Rombauts, S.; Rouze, P.; Worden, A. Z.; Robbens, S.; Partensky, F.; Degroeve, S.; Echeynie, S.; Cooke, R.; Saeys, Y.; Wuyts, J.; Jabbari, K.; Bowler, C.; Panaud, O.; Piegu, B.; Ball, S. G.; Ral, J.-P.; Bouget, F.-Y.; Piganeau, G.; De Baets, B.; Picard, A.; Delseny, M.; Demaille, J.; Van De Peer, Y.; Moreau, H. (2006). "Genome analysis of the smallest free-living eukaryote Ostreococcus tauri unveils many unique features". Proceedings of the National Academy of Sciences. 103 (31): 11647–11652. Bibcode:2006PNAS..10311647D. doi:10.1073/pnas.0604795103. PMC 1544224. PMID 16868079.
- ^ O'Kelly, Charles J.; Sieracki, Michael E.; Thier, Edward C.; Hobson, Ilana C. (2003). "A Transient Bloom of Ostreococcus (Chlorophyta, Prasinophyceae) in West Neck Bay, Long Island, New York". Journal of Phycology. 39 (5): 850–854. Bibcode:2003JPcgy..39..850O. doi:10.1046/j.1529-8817.2003.02201.x. S2CID 84541779.
- ^ Yau, Sheree; Hemon, Claire; Derelle, Evelyne; Moreau, Hervé; Piganeau, Gwenaël; Grimsley, Nigel (2016). "A Viral Immunity Chromosome in the Marine Picoeukaryote, Ostreococcus tauri". PLOS Pathogens. 12 (10): e1005965. doi:10.1371/journal.ppat.1005965. PMC 5082852. PMID 27788272.
- ^ Weitz, Joshua S.; Wilheilm, Steven W. (1 July 2013). "An Ocean of Viruses". The Scientist.
- ^ Worden, A. Z.; Lee, J.-H.; Mock, T.; Rouze, P.; Simmons, M. P.; Aerts, A. L.; Allen, A. E.; Cuvelier, M. L.; Derelle, E.; Everett, M. V.; Foulon, E.; Grimwood, J.; Gundlach, H.; Henrissat, B.; Napoli, C.; McDonald, S. M.; Parker, M. S.; Rombauts, S.; Salamov, A.; von Dassow, P.; Badger, J. H.; Coutinho, P. M.; Demir, E.; Dubchak, I.; Gentemann, C.; Eikrem, W.; Gready, J. E.; John, U.; Lanier, W.; et al. (2009). "Green Evolution and Dynamic Adaptations Revealed by Genomes of the Marine Picoeukaryotes Micromonas". Science. 324 (5924): 268–272. Bibcode:2009Sci...324..268W. doi:10.1126/science.1167222. PMID 19359590. S2CID 206516961.
- ^ Evans, C.; Archer, SD; Jacquet, S.; Wilson, WH (2003). "Direct estimates of the contribution of viral lysis and microzooplankton grazing to the decline of a Micromonas spp. Population". Aquatic Microbial Ecology. 30: 207–219. doi:10.3354/ame030207.
Further reading
[edit]- Van Etten, James L. (2003). "Unusual Life Style of Giant Chlorella Viruses". Annual Review of Genetics. 37: 153–195. doi:10.1146/annurev.genet.37.110801.143915. PMID 14616059.
- Van Etten, James L.; Meints, Russel H. (1999). "Giant Viruses Infecting Algae". Annual Review of Microbiology. 53: 447–494. doi:10.1146/annurev.micro.53.1.447. PMID 10547698. S2CID 7346326.
- Iyer, Lakshminarayan M.; Balaji, S.; Koonin, Eugene V.; Aravind, L. (2006). "Evolutionary genomics of nucleo-cytoplasmic large DNA viruses". Virus Research. 117 (1): 156–184. doi:10.1016/j.virusres.2006.01.009. PMID 16494962.
- Raoult, D.; Audic, Stéphane; Robert, Catherine; Abergel, Chantal; Renesto, Patricia; Ogata, Hiroyuki; La Scola, Bernard; Suzan, Marie; Claverie, Jean-Michel (2004). "The 1.2-Megabase Genome Sequence of Mimivirus". Science. 306 (5700): 1344–1350. Bibcode:2004Sci...306.1344R. doi:10.1126/science.1101485. PMID 15486256. S2CID 84298461.
- World of Chlorella Viruses Home Page
